
CytoDataFrame#
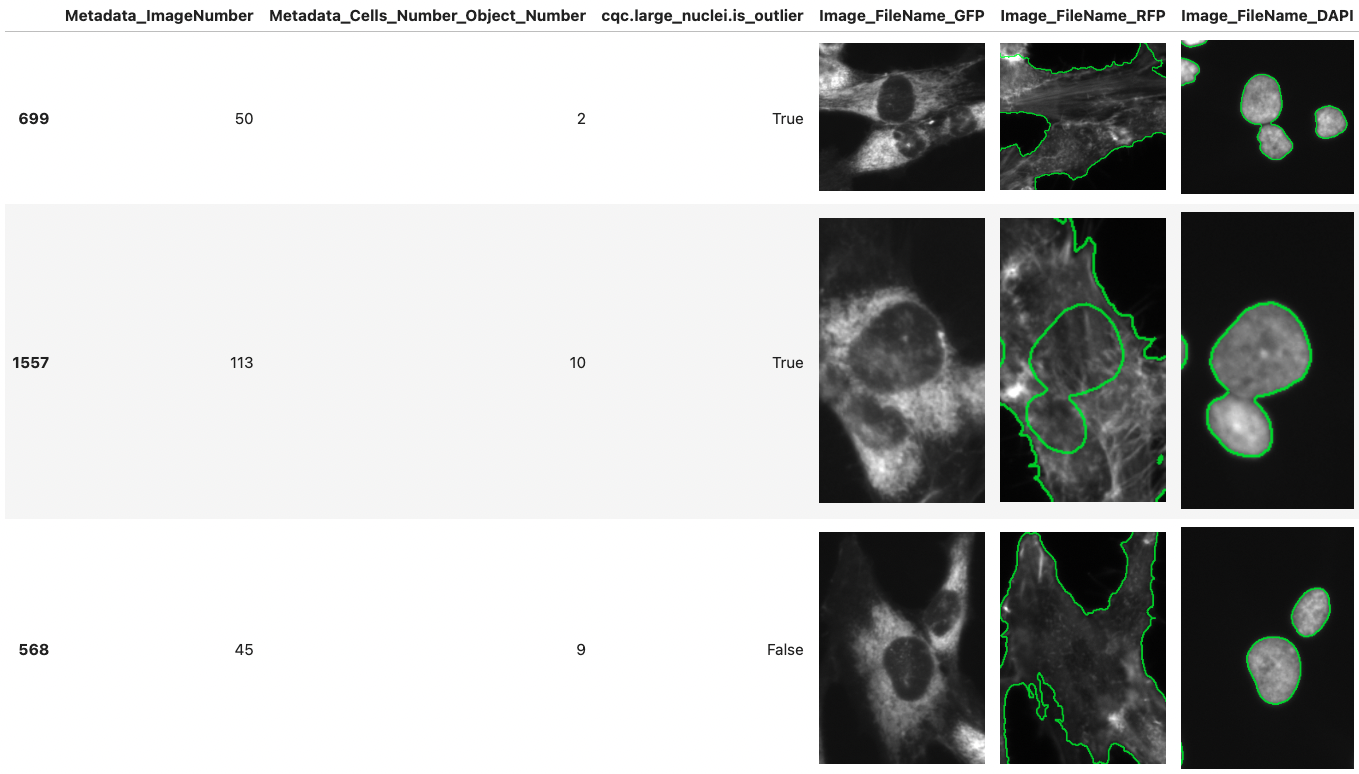 CytoDataFrame extends Pandas functionality to help display single-cell profile data alongside related images.
CytoDataFrame extends Pandas functionality to help display single-cell profile data alongside related images.
CytoDataFrame is an advanced in-memory data analysis format designed for single-cell profiling, integrating not only the data profiles but also their corresponding microscopy images and segmentation masks. Traditional single-cell profiling often excludes the associated images from analysis, limiting the scope of research. CytoDataFrame bridges this gap, offering a purpose-built solution for comprehensive analysis that incorporates both the data and images, empowering more detailed and visual insights in single-cell research.
CytoDataFrame is best suited for work within Jupyter notebooks. With CytoDataFrame you can:
View image objects alongside their feature data using a Pandas DataFrame-like interface.
Highlight image objects using mask or outline files to understand their segmentation.
Adjust image displays on-the-fly using interactive slider widgets.
Automatically detect 3D image volumes and render interactive trame views in notebooks when 3D dependencies are installed (with graceful fallback otherwise).
For 3D notebook display behavior:
3D-aware rendering is enabled by default (
display_options={"auto_trame_for_3d": True}).Disable automatic trame switching with
display_options={"auto_trame_for_3d": False}.Force trame layout regardless of auto-detection with
display_options={"view": "trame"}.
For row display in notebook/widget tables:
CytoDataFrame respects pandas display settings (
display.max_rows,display.min_rows).When the table is larger than
display.max_rows, the widget table inserts a midpoint ellipsis row (…) to indicate omitted rows.You can control truncation behavior by changing pandas display options before rendering.
📓 Want to see CytoDataFrame in action? Check out our example notebook for a quick tour of its key features.
✨ CytoDataFrame development began within coSMicQC - a single-cell profile quality control package. Please check out our work there as well!
Installation#
Install CytoDataFrame from source using the following:
# install from pypi
pip install cytodataframe
# or install directly from source
pip install git+https://github.com/cytomining/CytoDataFrame.git
Contributing, Development, and Testing#
Please see our contributing documentation for more details on contributions, development, and testing.



